CASE STUDIES
Transforming our ability to understand and manage our environment.
Examples of the Genomics work supported by NEOF and the Liverpool and Sheffield nodes of the NERC Biomolecular Analysis Facility (the predecessor to NEOF) from 2019.
Examples of Metabolomics work supported by the Centre for Proteomics Research, University of Liverpool.
Examples of Proteomics work supported by the Centre for Proteomics Research, University of Liverpool.


Single-cell consequences of X-linked meiotic drive in stalk-eyed flies (2025)
NEOF supported single-cell RNA sequencing.
Price et al (2025) PLOS Genetics. 10.1371/journal.pgen.1011816
Invisible
READ MORE
Meiotic drivers, a class of selfish gene, are frequently located on sex chromosomes and have dramatic impacts on gamete development. However, our understanding of their molecular consequences for gamete production and sex chromosome regulation has focused on a handful of model organisms. In this study, we use single-cell RNA-sequencing approaches to produce a single-cell atlas of the testis of the stalk-eyed fly, Teleopsis dalmanni. This species harbours an X-linked meiotic driver where drive males produce more than 90% female offspring. First, we generate a comprehensive profile of the cellular and transcriptional landscape of spermatogenesis. We show limited evidence for meiotic sex chromosome inactivation and unique patterns of dosage compensation across spermatogenesis, relative to both other dipterans and insects in general. Finally, by comparing single-cell expression data between standard and drive males, we show that although there are significant differences in genome regulation, broad expression dynamics in the testis are conserved in the presence of meiotic drive. Notably, we highlight key genes with perturbed expression as a potential consequence of the disruption of spermatogenesis by the X-linked meiotic driver.
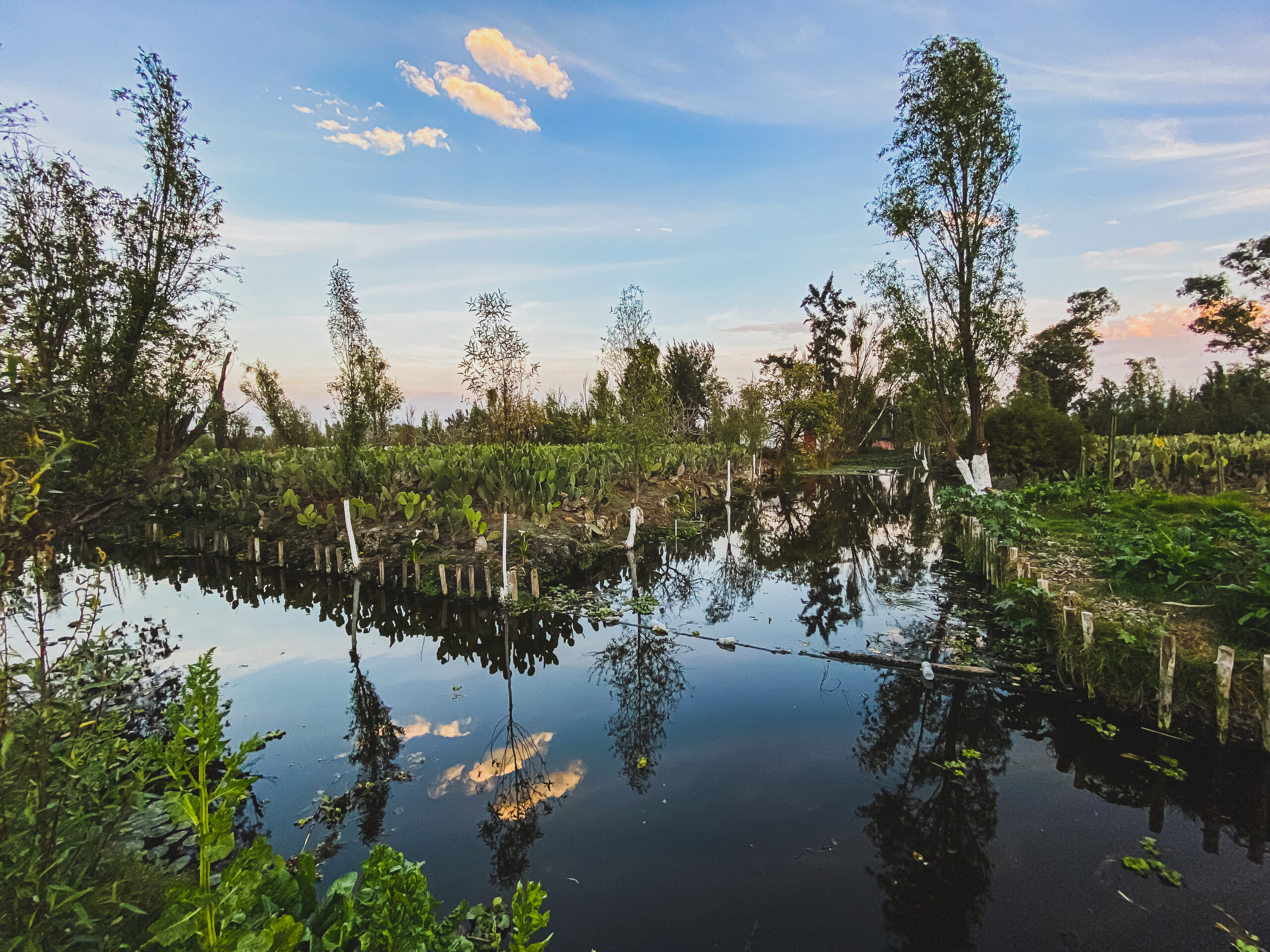

Persisting at the Edge of Ecological Collapse: The Impact of Urbanization on Fish and Amphibian Communities From Lake Xochimilco (2025)
NEOF supported a PhD student (Alejandro Maeda-Obregón) at the Visitor Facility, who used targeted and metabarcoding eDNA approaches to study the biodiversity of Lake Xochimilco in Mexico City, a World Heritage site and historic home of the critically endangered Axolotl salamander and other unique animals such as the Mexclapique live-bearing fish.
Maeda-Obregón et al (2025) Environmental DNA (10.1002/edn3.70147).
Invisible
READ MORE
Freshwater ecosystems are globally threatened by habitat loss, pollution, and invasive species, all of which are particularly acute in urban areas. To assess the impacts of urbanization on freshwater biodiversity—specifically the effects of alien species on native primary aquatic vertebrates—we investigated the World Heritage Site, Lake Xochimilco in Mexico City. Focusing on fishes and amphibians, we applied environmental DNA metabarcoding using primer pairs targeting mitochondrial 12S and 16S across the remnant lake and collected 14 aquatic environmental variables for sampled sites. Our survey recovered ca. 60% of Lake Xochimilco’s historically recorded fish and amphibian species, including rare species and novel taxa not detected by past traditional surveys. However, our findings imply a severely degraded wetland, with alpha diversity indices indicating a low-diversity ecosystem dominated by alien fishes. Beta diversity analysis revealed a heterogeneous ecosystem that may be driven partially by the presence of alien fish, particularly cyprinids. Environmental variables linked to pollution predicted the presence of non-native fish families. We also found evidence that some species prefer to occupy different water bodies within the lake remnant. Despite the ongoing degradation of this ecosystem, native and endemic fauna are persisting, although detections were typically rare. We found no evidence of the Critically Endangered axolotl salamanders (Ambystoma sp.) from wild sites; however, we detected their presence in one wildlife refuge, highlighting the potential of refuges to prevent complete extinction in the wild. We also found evidence of cryptic taxonomic diversity in Lithobates frogs and evidence of endemic genera, including the threatened mexclapique fish (Girardinicthys viviparus). These fishes are considered extirpated, suggesting remnant populations persist undetected by traditional surveys. Despite clear evidence of an ecosystem under extreme decline compared to historical biological records, our study demonstrates the potential for restoration, given the presence of native freshwater species and the success of wildlife refuges.
, , , et al. 2025. “ Persisting at the Edge of Ecological Collapse: The Impact of Urbanization on Fish and Amphibian Communities From Lake Xochimilco.” Environmental DNA 7, no. 4: e70147. https://doi.org/10.1002/edn3.70147.
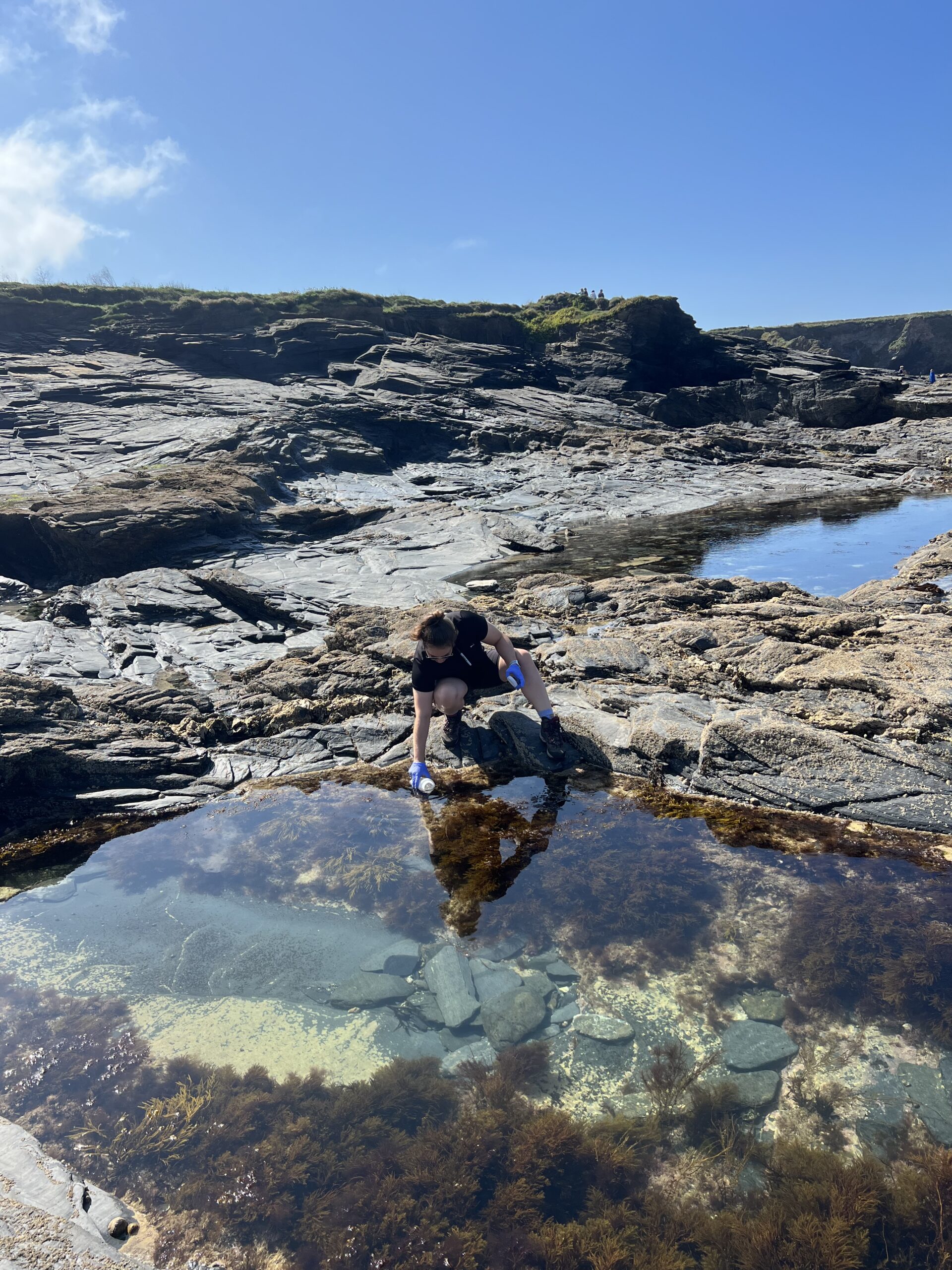

Characterizing Rocky Intertidal Biodiversity Using Environmental DNA Metabarcoding From Local to National Scales (2025)
NEOF supported a PhD student (Dina Simons) at the Visitor Facility investigating metabarcoding of intertidal eDNA, including from rockpools, as a tool to monitor both local and national marine biodiversity across the UK.
Simons et al (2025) Environmental DNA (10.1002/edn3.70203).
Invisible
READ MORE
Efficient and scalable methods for monitoring marine biodiversity are critical for understanding ecological change in coastal environments, given the limited resources available. Environmental DNA (eDNA) metabarcoding shows promise for monitoring coastal taxa, but its ability to differentiate communities from different locations remains insufficiently understood, particularly in dynamic marine environments. Here, we evaluate the effectiveness and resolution capacity of eDNA metabarcoding in detecting rocky intertidal taxa across three spatial scales—national, regional, and local—in the United Kingdom. Onshore surface-water samples were collected from 32 sites across five UK Regional Seas from rockpools in high and low shore zones, as well as directly from the sea. We detected 1026 target taxa within 442 families and 19 phyla using two established markers targeting invertebrates (COI) and macroalgae (18S). Distinct eDNA signals were found at all spatial scales, indicating local discreteness even between vertical shore heights within the same sites. Communities were more discrete at larger scales (i.e., between regions) than at smaller scales (i.e., between shore heights). eDNA signals were more strongly structured by geographical location than by vertical shore height as a probable consequence of greater DNA homogenization over the tidal cycle at smaller spatial scales. Established ecological zonation patterns were reflected in eDNA signals, with higher richness at lower shore heights, reflecting abiotic stress gradients. Detections of cold-affinity boreal species increased with latitude, while warm-affinity lusitanian species declined with latitude. Our work supports the utility of eDNA metabarcoding for multiscale biodiversity monitoring in dynamic marine environments and for detections beyond this study’s target taxa. We recommend the adoption of scale-appropriate sampling protocols to optimize the benefits of eDNA, such as prioritizing open water sampling at high tide for broad-scale assessments and rockpool sampling at low tide for capturing local-scale patterns. Future work should validate detections through direct visual comparisons.
, , , , , and (2025) Characterizing Rocky Intertidal Biodiversity Using Environmental DNA Metabarcoding From Local to National Scales. Environmental DNA 7, no. 5: e70203. https://doi.org/10.1002/edn3.70203.
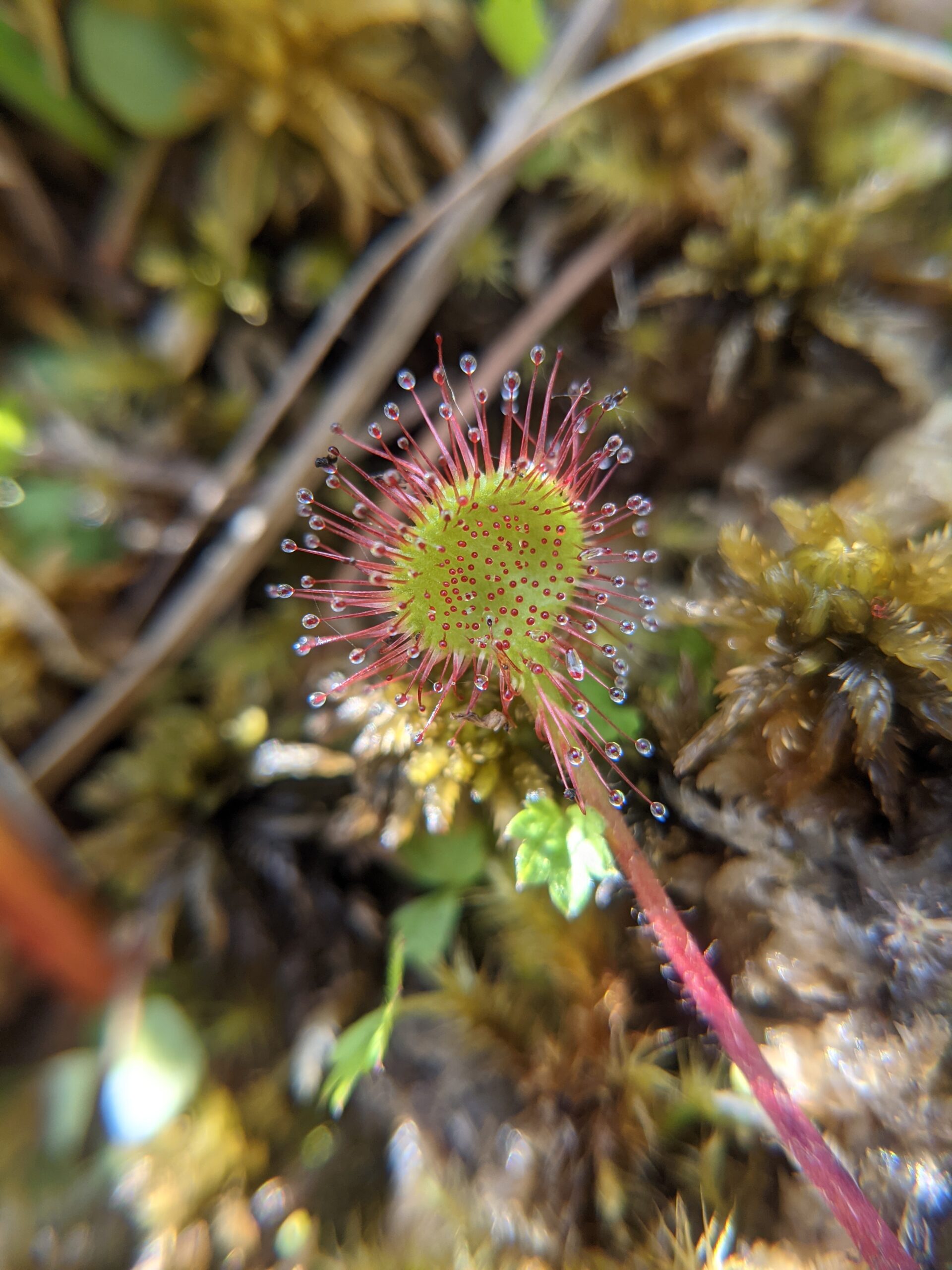

An acidophilic fungus promotes prey digestion in a carnivorous plant (2024)
NEOF supported a PhD student (Brandon Shaw) who performed metabarcoding at the Visitor Facility to reveal that a fungus (Acrodontium crateriforme) helps digest prey in a carnivorous plant (Drosera spatulata). Fungal endophyte community composition data showed that A. crateriforme is present and often dominant in the communities of three other Drosera species in the UK and US. This provides evidence of a likely global, ancient, and important symbiosis for this plant, which shifts the perspective on how plant carnivory can evolve.
Sun PF et al (2024) Nature Microbiology (10.1038/s41564-024-01766-y)
Invisible
READ MORE
Leaves of the carnivorous sundew plants (Drosera spp.) secrete mucilage that hosts microorganisms, but whether this microbiota contributes to prey digestion is unclear. We identified the acidophilic fungus Acrodontium crateriforme as the dominant species in the mucilage microbial communities, thriving in multiple sundew species across the global range. The fungus grows and sporulates on sundew glands as its preferred acidic environment, and its presence in traps increased the prey digestion process. A. crateriforme has a reduced genome similar to other symbiotic fungi. During A. crateriforme–Drosera spatulata coexistence and digestion of prey insects, transcriptomes revealed significant gene co-option in both partners. Holobiont expression patterns during prey digestion further revealed synergistic effects in several gene families including fungal aspartic and sedolisin peptidases, facilitating prey digestion in leaves, as well as nutrient assimilation and jasmonate signalling pathway expression. This study establishes that botanical carnivory is defined by adaptations involving microbial partners and interspecies interactions.

Competing adaptations maintain nonadaptive variation in a wild cricket population (2024)
NEOF generated genomics and transcriptomics data for this study.
Rayner et al (2024), Proceedings of the National Academy of Sciences (10.1073/pnas.2317879121)
Invisible
READ MORE
Field crickets in Hawaii have largely stopped singing. Crickets sing by rubbing their wings together and a parasitoid wasp recently introduced to the islands zeroes in this song to lay its eggs in the cricket. This has led to extremely strong selection for mutant wing morphologies that make crickets unable to sing. Here Rayner et al have used genomics and transcriptomics to elucidate the genetic basis of different mutations that have appeared and find that the co-occurrence of different mutations prevents the spread of any single mutation to fixation. The work presents a remarkable illustration of rapid evolution in response to a sudden change in the environment. It was conducted by an international team of UK and US researchers.

Admixture and reproductive skew shape the conservation value of ex situ populations of the Critically Endangered eastern black rhino (2024)
The work was supported by student access to the NEOF Visitor Facility.
Elsner-Gearing et al (2024), Conservation Genetics. (10.1007/s10592-024-01611-z)
Invisible
READ MORE
The black rhino is critically endangered. Poaching has caused a drastic drop in numbers from ~850,000 to ~6,000 today, the fragmentation of the remaining population into many sub-populations and consequent loss of genetic diversity. Conservation management to protect genetic diversity is therefore important. Here Elsner-Gearing et al used molecular markers to reconstruct black rhino pedigrees in zoos and wild populations. They showed in both cases that male biased reproductive skew was prevalent, which meant that one male could father many offspring, effectively reducing the genetic diversity in a population. However, its effect on loss of genetic diversity was mitigated in breeding strategies used in zoos. The work shows how molecular methods can support conservation efforts in zoos and in situ. The work was supported by student access to the visitor facility, and involved collaboration between UK and international researchers and between academia and conservation NGOs.

Soil microbiomes show consistent and predictable responses to extreme events (2024)
Meta-genomic sequencing was conducted at NEOF.
Knight CG et al (2024) Nature (10.1038/s41586-024-08185-3)
Invisible
READ MORE
Microbial communities (microbiomes) within the soil are essential to the health and functioning of entire ecosystems. Here Knight et al used genomic methods to study how soil microbiomes from 30 grasslands across Europe respond to extreme events such as heat, drought and flooding, which may all become more frequent with climate change. They showed that soil microbiomes from different sites exhibited repeatable gene- and species-level changes in response to extreme events. Their findings provide a basis to make predictions about the effect of extreme climatic events on soil ecosystem function. Meta-genomic sequencing was conducted at NEOF. The work was led by the University of Manchester and involved researchers from across Europe.

Effects of an IgE receptor polymorphism acting on immunity, susceptibility to infection, and reproduction in a wild rodent (2023)
NEOF genomics made this non-model species accessible to investigation.
Wanelik et al (2023) eLife 10.7554/elife.77666
Invisible
READ MORE
How variation between individuals affects their ability to deal with infection is a fundamental question that is difficult to address in laboratory studies that lack the genetic and environmental variation found in natural populations. The field vole (Microtus agrestis) is related to the laboratory mouse but exposed to the natural environment. Here NEOF genomics made this non-model species accessible to investigation. Wanelik et al were able to discover genetic polymorphisms that affected expression of immunological genes, analyse these within a gene regulatory network and examine their consequent effects for susceptibility to infection and to reproduction.

Life on Earth can grow on extraterrestrial organic carbon (2023)
Waajen et al (2023) Scientific Reports, 10.1038/s41598-024-54195-6
Invisible
READ MORE
The universe is full of abiotic carbon, found in meteorites, that could serve as a carbon source for life on other planets or the early Earth. Here researchers from NEOF supported a 13C-stable isotope labelling experiment in combination with O-PITR spectroscopy to show that carbon can be transferred from meteoric carbon to microbial biomass. This supports the potential for a heterotrophic metabolism in early living systems.

Gut microbiota of the critically endangered Saiga antelope across two wild populations in a year without mass mortality (2023)
NEOF supported metagenetic sequencing.
Hanski et al (2023) Scientific Reports, 10.1038/s41598-023-44393-z
Invisible
READ MORE
Saiga antelope are a long-distance migratory species, predominantly found in Kazakhstan. They are critically endangered and have been subject to mass mortality events, which are putatively associated with the presence of Pasteurella. Saiga are wary of human contact but their faeces can be sampled. Here Hanski and colleagues interrogated the gut microbiome found in faeces to provide baseline data for species composition in Saiga and compared this to other antelope. In healthy populations, no Pasteurella was found. While further validation is required, this provides the possibility to develop methods to non-invasively sample this endangered population to detect pathogen species. NEOF supported metagenetic sequencing.

Characterising a genetic stronghold amidst pervasive admixture: Morelet’s crocodiles (Crocodylus moreletii) in central Yucatan (2023)
NEOF supported a PhD student (José Barão-Nóbrega) at the Visitor Facility.
Barão-Nóbrega et al (2023) Conservation Genetics, 10.1007/s10592-023-01544-z
Invisible
READ MORE
Hybridisation presents a threat to endangered species. Populations of Morelet’s crocodiles (Crocodylus moreletii) in Mexico,are threatened by interbreeding with the more numerous American crocodile (C. acutus). Here, molecular genotyping was used to investigate genome-wide hybridisation and introgression across the C. moreletii range. This discovered a stronghold of populations that were relatively resistant to hybridisation. This provides information that can be used to guide conservation of this species.
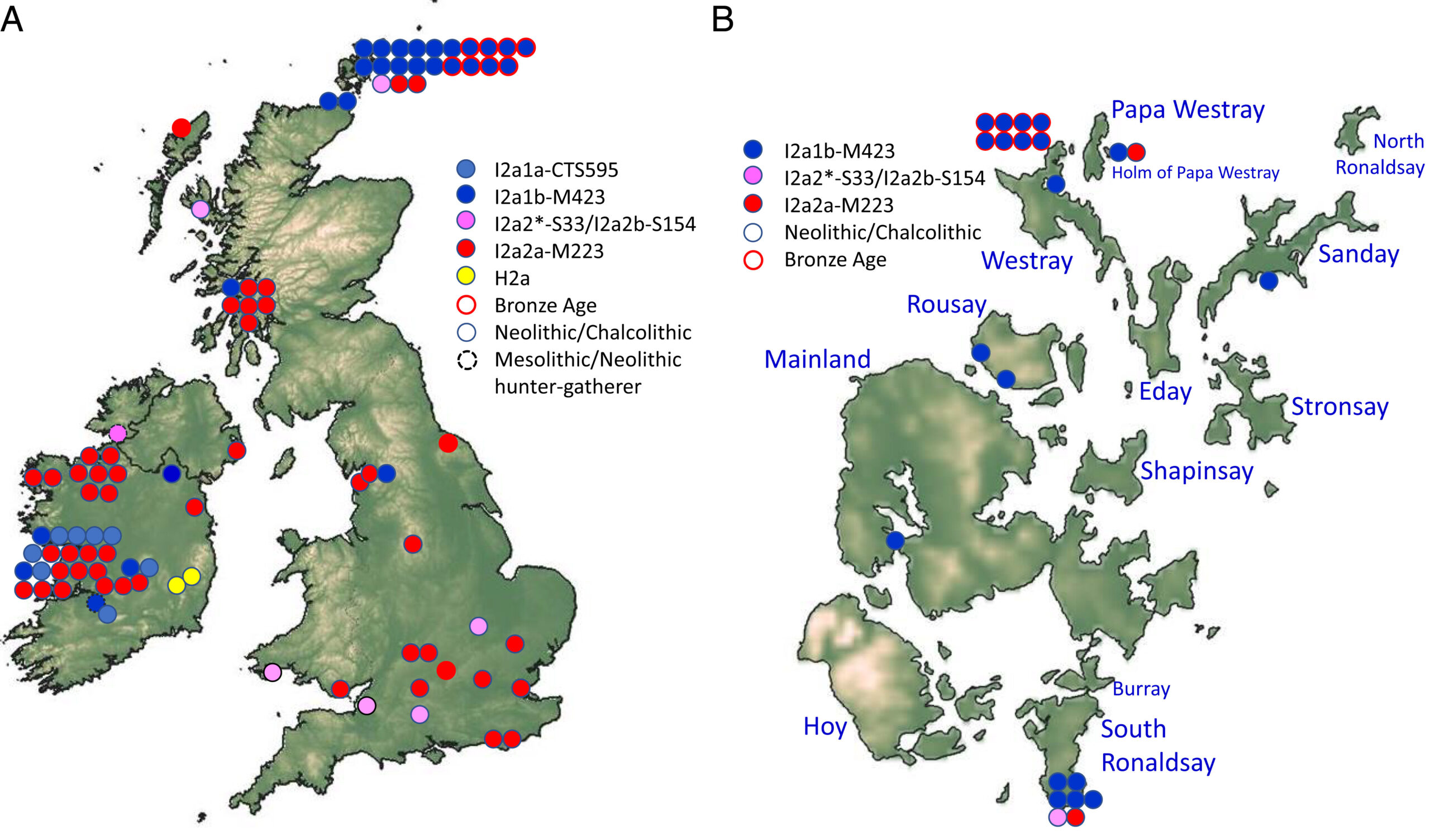
Ancient DNA at the edge of the world: Continental immigration and the persistence of Neolithic male lineages in Bronze Age Orkney (2022)
Genome sequencing by NEOF helped to show that Orkney in the Bronze Age showed far more immigration, and hence movement of both people and culture, than previously believed. It was particularly striking that immigration was almost solely of females, with male Y lineages persisting for at least a thousand years.
Dulias et al (2022) PNAS 10.1073/pnas.2108001119
Invisible
READ MORE
Orkney was a major cultural center during the Neolithic, 3800 to 2500 BC. Farming flourished, permanent stone settlements and chambered tombs were constructed, and long-range contacts were sustained. From ∼3200 BC, the number, density, and extravagance of settlements increased, and new ceremonial monuments and ceramic styles, possibly originating in Orkney, spread across Britain and Ireland. By ∼2800 BC, this phenomenon was waning, although Neolithic traditions persisted to at least 2500 BC. Unlike elsewhere in Britain, there is little material evidence to suggest a Beaker presence, suggesting that Orkney may have developed along an insular trajectory during the second millennium BC. We tested this by comparing new genomic evidence from 22 Bronze Age and 3 Iron Age burials in northwest Orkney with Neolithic burials from across the archipelago. We identified signals of inward migration on a scale unsuspected from the archaeological record: As elsewhere in Bronze Age Britain, much of the population displayed significant genome-wide ancestry deriving ultimately from the Pontic-Caspian Steppe. However, uniquely in northern and central Europe, most of the male lineages were inherited from the local Neolithic. This suggests that some male descendants of Neolithic Orkney may have remained distinct well into the Bronze Age, although there are signs that this had dwindled by the Iron Age. Furthermore, although the majority of mitochondrial DNA lineages evidently arrived afresh with the Bronze Age, we also find evidence for continuity in the female line of descent from Mesolithic Britain into the Bronze Age and even to the present day.
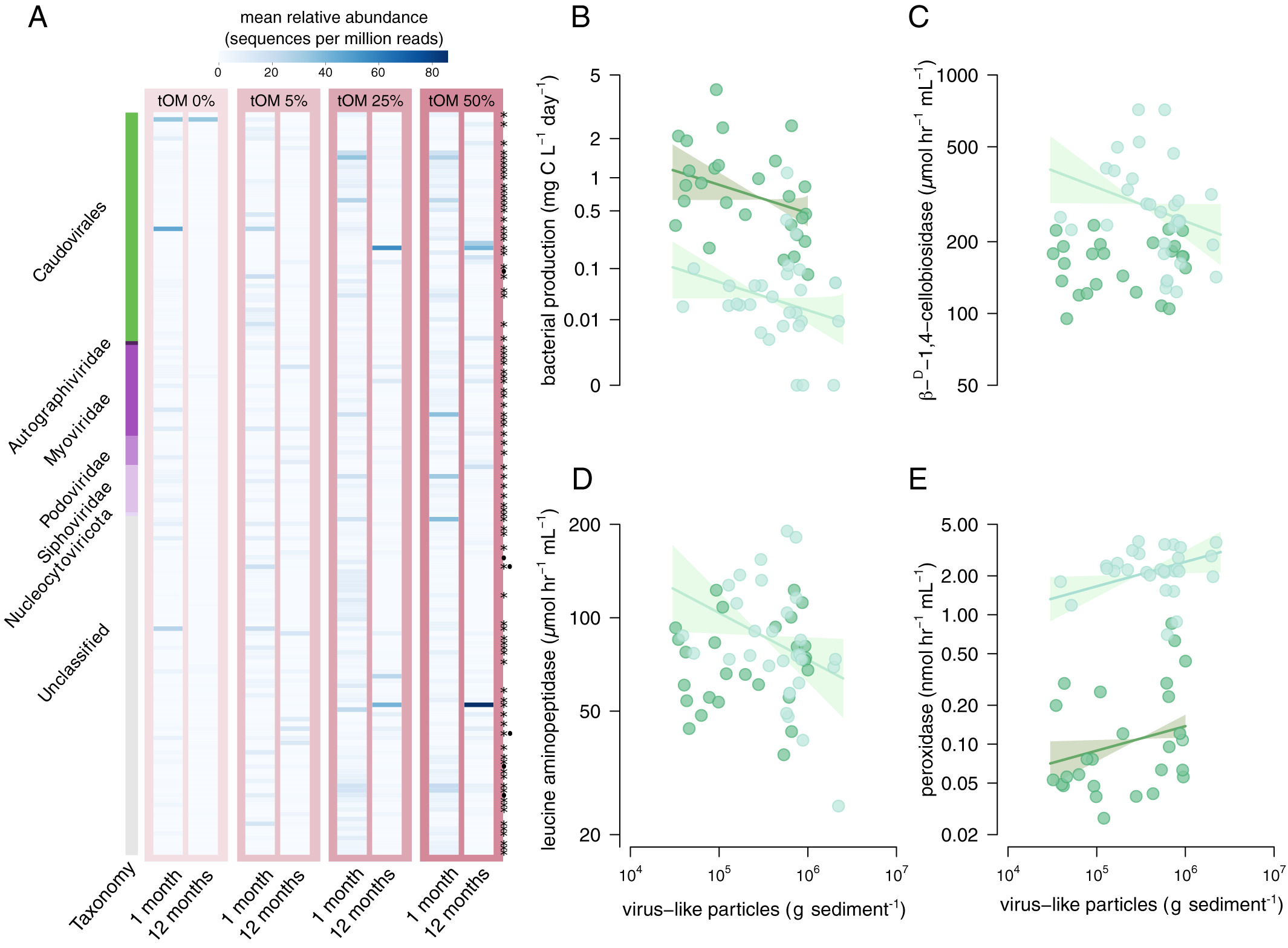
Viruses direct carbon cycling in lake sediments under global change (2022)
eDNA analysis at NEOF helped to show the importance of viruses to carbon cycling in lake sediments and to identify viral genes that affect the processing of carbon compounds. These results argue that viruses need to be included in any predictive model of the freshwater carbon cycle in response to climate change.
(Braga et al 2022 PNAS)
Invisible
READ MORE
Global change is altering the vast amount of carbon cycled by microbes between land and freshwater, but how viruses mediate this process is poorly understood. Here, we show that viruses direct carbon cycling in lake sediments, and these impacts intensify with future changes in water clarity and terrestrial organic matter (tOM) inputs. Using experimental tOM gradients within sediments of a clear and a dark boreal lake, we identified 156 viral operational taxonomic units (vOTUs), of which 21% strongly increased with abundances of key bacteria and archaea, identified via metagenome-assembled genomes (MAGs). MAGs included the most abundant prokaryotes, which were themselves associated with dissolved organic matter (DOM) composition and greenhouse gas (GHG) concentrations. Increased abundances of virus-like particles were separately associated with reduced bacterial metabolism and with shifts in DOM toward amino sugars, likely released by cell lysis rather than higher molecular mass compounds accumulating from reduced tOM degradation. An additional 9.6% of vOTUs harbored auxiliary metabolic genes associated with DOM and GHGs. Taken together, these different effects on host dynamics and metabolism can explain why abundances of vOTUs rather than MAGs were better overall predictors of carbon cycling. Future increases in tOM quantity, but not quality, will change viral composition and function with consequences for DOM pools. Given their importance, viruses must now be explicitly considered in efforts to understand and predict the freshwater carbon cycle and its future under global environmental change.
Braga et al (2022) Viruses direct carbon cycling in lake sediments under global change. PNAS.
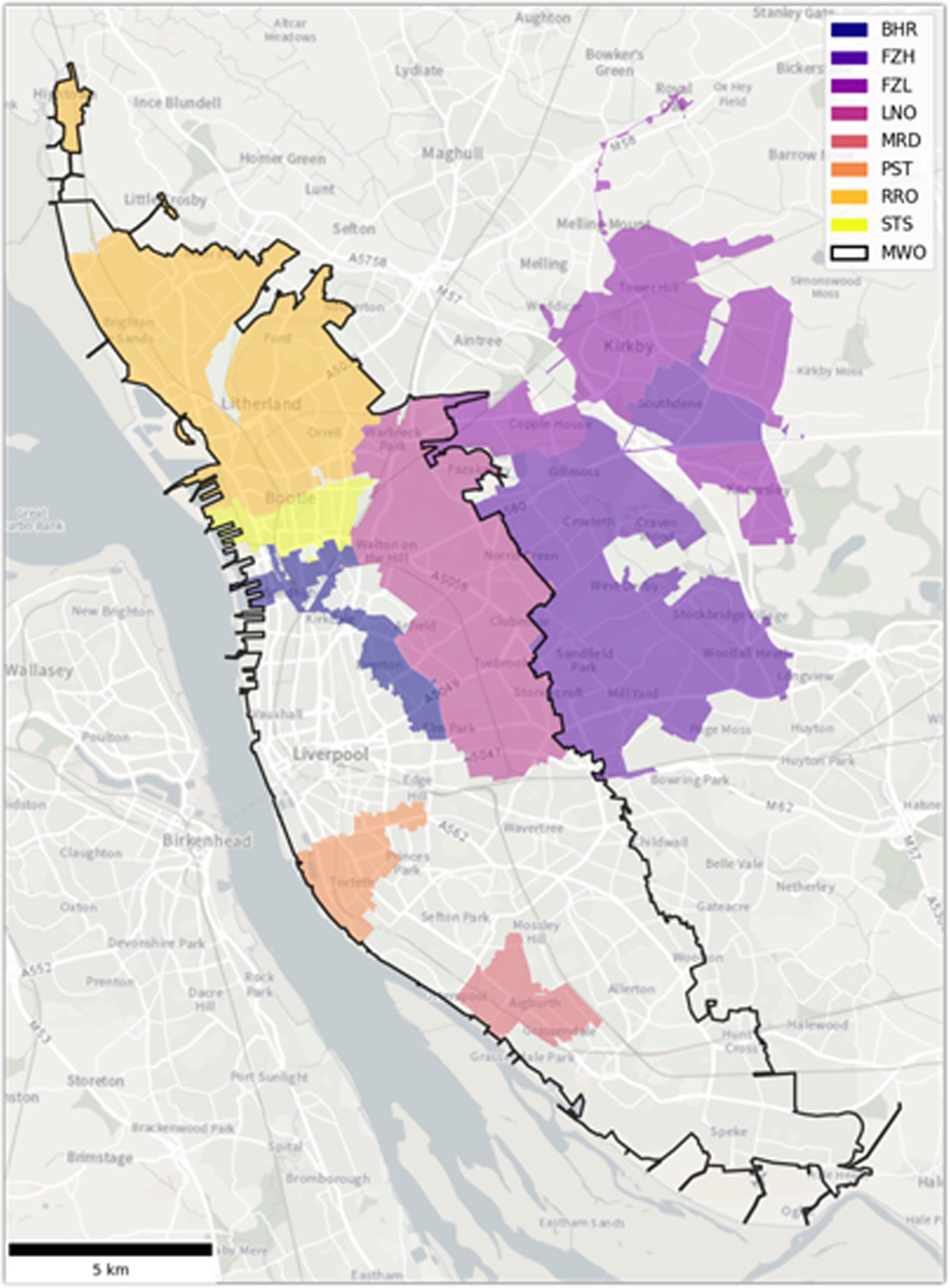
City-wide wastewater genomic surveillance through the successive emergence of SARS-CoV-2 Alpha and Delta variants (2022)
NEOF expertise and capability was used to develop and implement wastewater based epidemiology to detect and monitor SARS-CoV-2 variants. Here we demonstrated this capability to detect the emergence and rise of Alpha and Delta variants within an urban sewage system that matched clinical sampling but at a fraction of the cost.
Brunner et al (2022) Water Research 226, 119306
Invisible
READ MORE
Genomic surveillance of SARS-CoV-2 has provided a critical evidence base for public health decisions throughout the pandemic. Sequencing data from clinical cases has helped to understand disease transmission and the spread of novel variants. Genomic wastewater surveillance can offer important, complementary information by providing frequency estimates of all variants circulating in a population without sampling biases. Here we show that genomic SARS-CoV-2 wastewater surveillance can detect fine-scale differences within urban centres, specifically within the city of Liverpool, UK, during the emergence of Alpha and Delta variants between November 2020 and June 2021. Furthermore, wastewater and clinical sequencing match well in the estimated timing of new variant rises and the first detection of a new variant in a given area may occur in either clinical or wastewater samples. The study’s main limitation was sample quality when infection prevalence was low in spring 2021, resulting in a lower resolution of the rise of the Delta variant compared to the rise of the Alpha variant in the previous winter. The correspondence between wastewater and clinical variant frequencies demonstrates the reliability of wastewater surveillance. However, discrepancies in the first detection of the Alpha variant between the two approaches highlight that wastewater monitoring can also capture missing information, possibly resulting from asymptomatic cases or communities less engaged with testing programmes, as found by a simultaneous surge testing effort across the city.
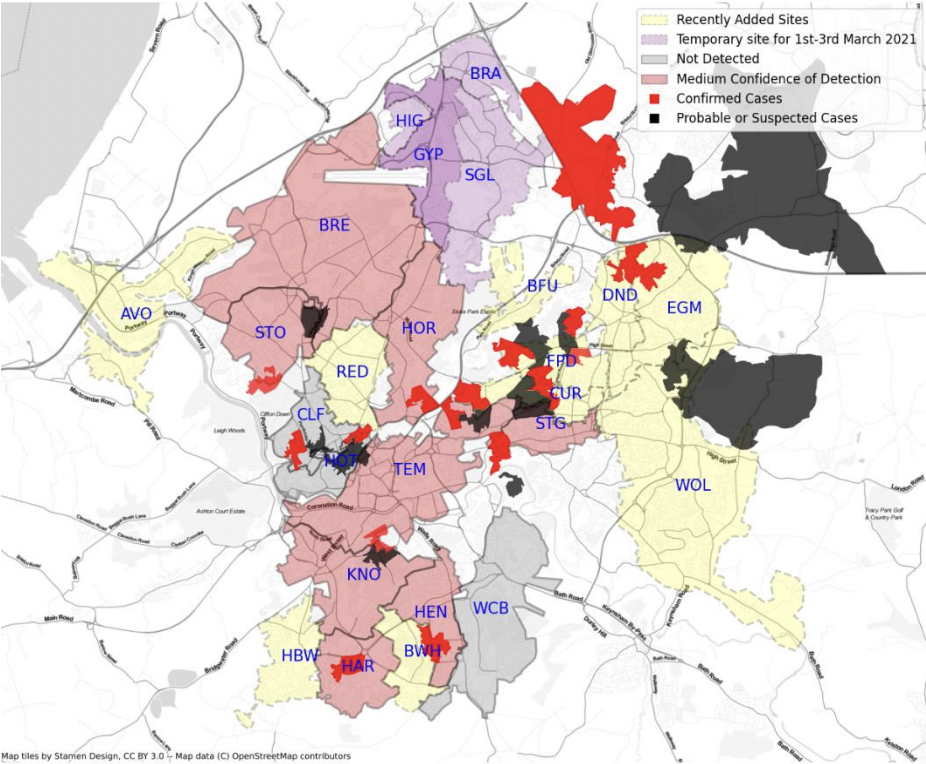
Environmental monitoring of SARS-CoV-2 (2021)
NEOF led the design of sequencing assays and generation data and analysis.
- Hilary et al (2021) Water Research 200, 117214
- Brown et al (2021) Summary for SAGE
Invisible
READ MORE
In support of the UK national response to COVID-19, NEOF prioritised work to use our skills in eDNA to detect and characterise coronavirus in wastewater. Working with the University of Bangor, we were able to demonstrate that SARS-CoV-2 could be genome-sequenced from wastewater taken from treatment plants, and were able to characterise its genetic diversity through the first wave of the pandemic. Having demonstrated the potential of the method, we then worked with UK Health Security Agency (UKHSA) Monitoring Health Protection Programme to deliver national surveillance of variants in wastewater for more than 1000 samples per week. This initial work by NEOF led to a major consortium of sequencing laboratories supported by funding from UKHSA. Highlights have included a rapid report to the CMO of the alpha (“Kent”) variant present in samples from London through November and December 2020, and an outbreak of an E484K mutation in Bristol used as a case study in a paper to SAGE.
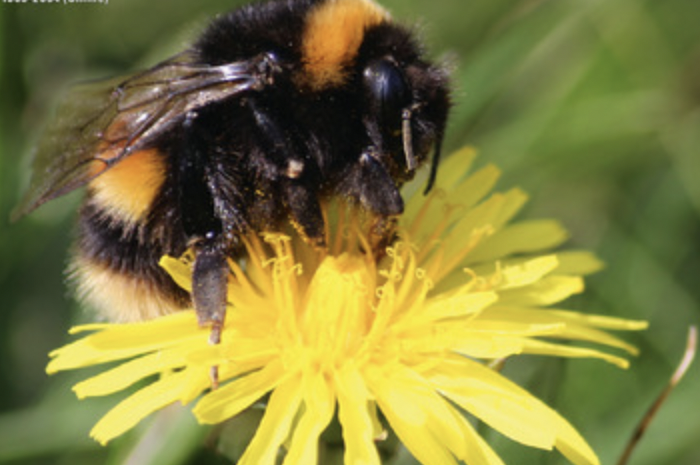

Bumblebee colony density on farmland (2021)
NEOF supported training in microsatellite genotyping and data analysis at the visitor facility. (Timberlake et al, 2021 JAE 1016-16)
Invisible
READ MORE
NEOF provided support and training to a PhD student project in the NEOF Visitor Facility to understand how the landscape affects the density of bumblebees, which are an important pollinator providing an essential ecosystem function. Molecular genotyping was used to estimate colony density on farmland and how this varied with nectar supply and garden cover. These results discovered the importance of the late spring as a bottleneck for resources, and highlighted the role that environmental stewardship programmes of farmland could have in increasing the density of bumblebee colonies, for example by modifying mowing regimes to delay flowering at field edges. This provides an example of how omics methods can provide an environmental solution within a systems approach. This work involved both international partners and conservation bodies, and received media interest, including in the farming press.
Timberlake et al. (2021) Bumblebee colony density on farmland is influenced by late-summer nectar supply and garden coverJ Appl Ecol 58, 1006–1016.
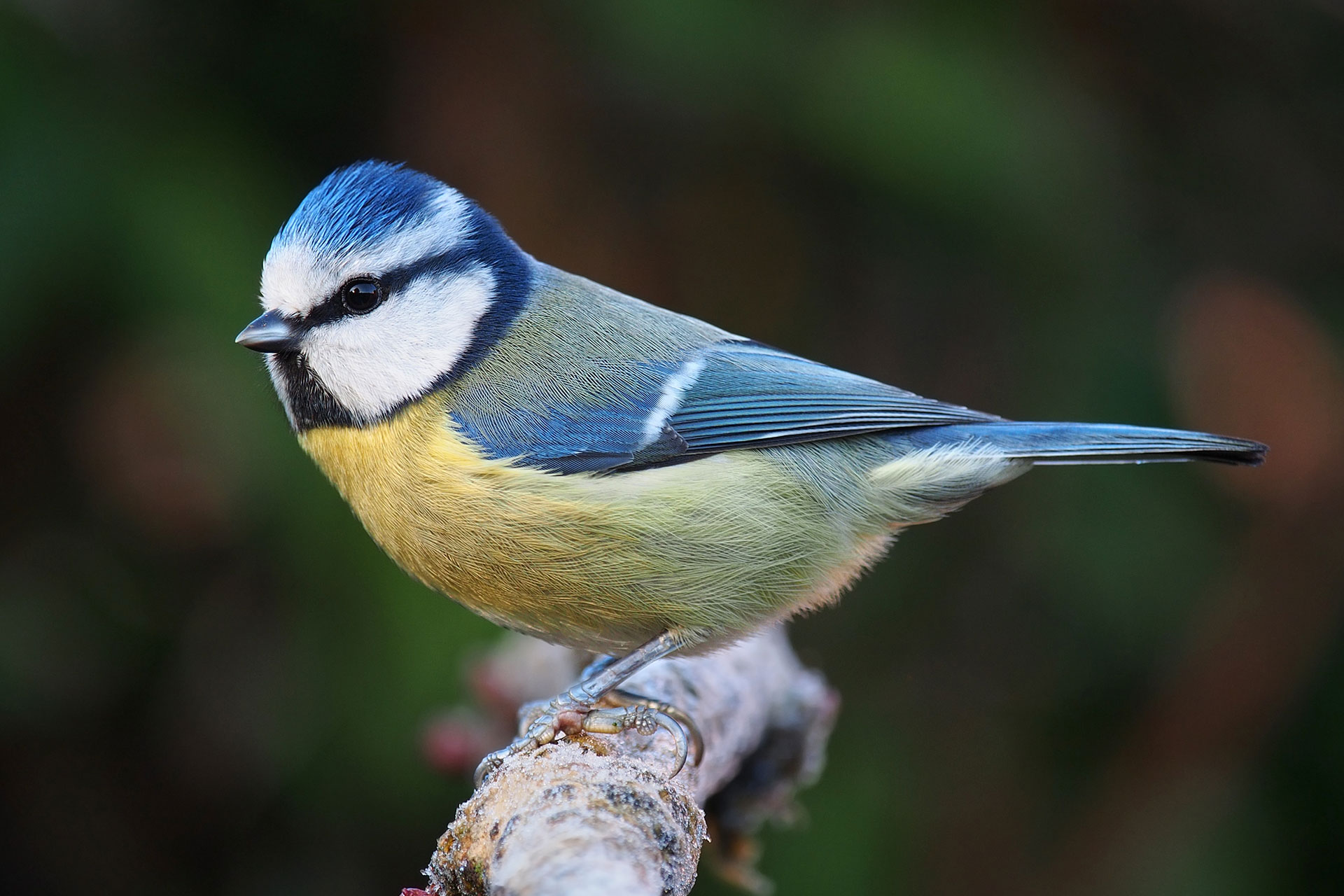

Gradients in richness and turnover of a forest passerine’s diet prior to breeding: A mixed model approach applied to faecal metabarcoding data (2020)
NBAF supported metabarcoding and bioinformatic analysis.
Invisible
READ MORE
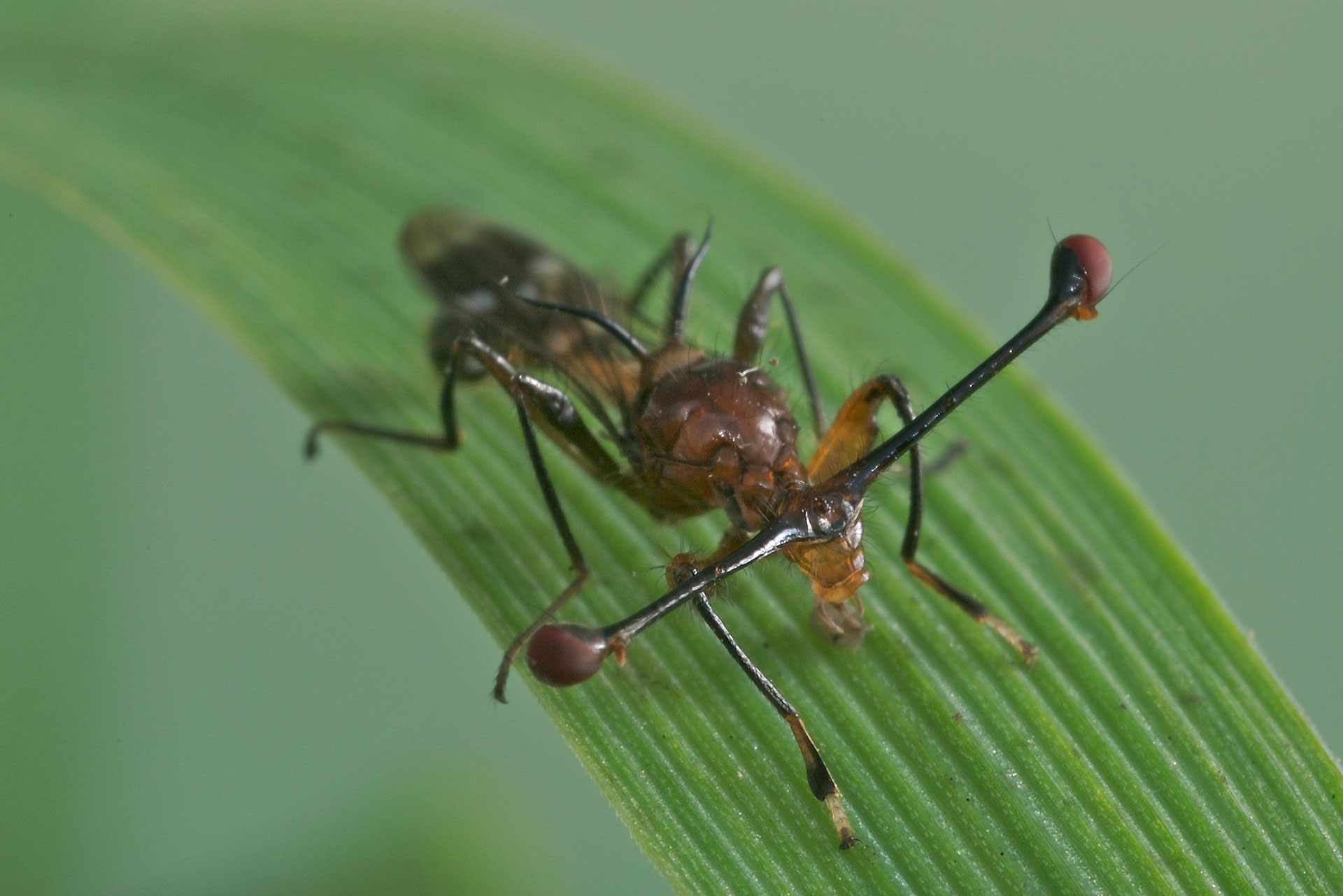

Meiotic drive reduces egg-to-adult viability in stalk-eyed flies (2019)
NBAF supported microsatellite genotyping and data analysis.
Invisible
READ MORE
Several species exhibit the phenomenon of Sex-Ratio (SR) meiotic drive, a selfish genetic element located on the X-chromosome that causes dysfunction of Y-bearing sperm, and one that has the potential to be exploited in the control of harmful insect vectors such as some mosquitoes. SR is transmitted to up to 100% of offspring, causing extreme sex-ratio bias. SR in several species is found in a stable polymorphism, suggesting that there must be strong frequency-dependent selection resisting its spread. This study investigated the effect of SR on female and male egg-to-adult viability in the Malaysian stalk-eyed fly, Teleopsis dalmanni. SR meiotic drive in this species is old, and appears to be broadly stable at a moderate (ca 20%) frequency. Large-scale controlled crosses were used to estimate the strength of selection acting against SR in female and male carriers. SR was found to reduce the egg-to-adult viability of both sexes. In females, homozygous females experienced greater reduction in viability and the deleterious effects of SR were additive. The male deficit in viability was not different from that in homozygous females. The study contributed to our understanding of how these reductions in egg-to-adult survival maintain the SR polymorphism in this species.
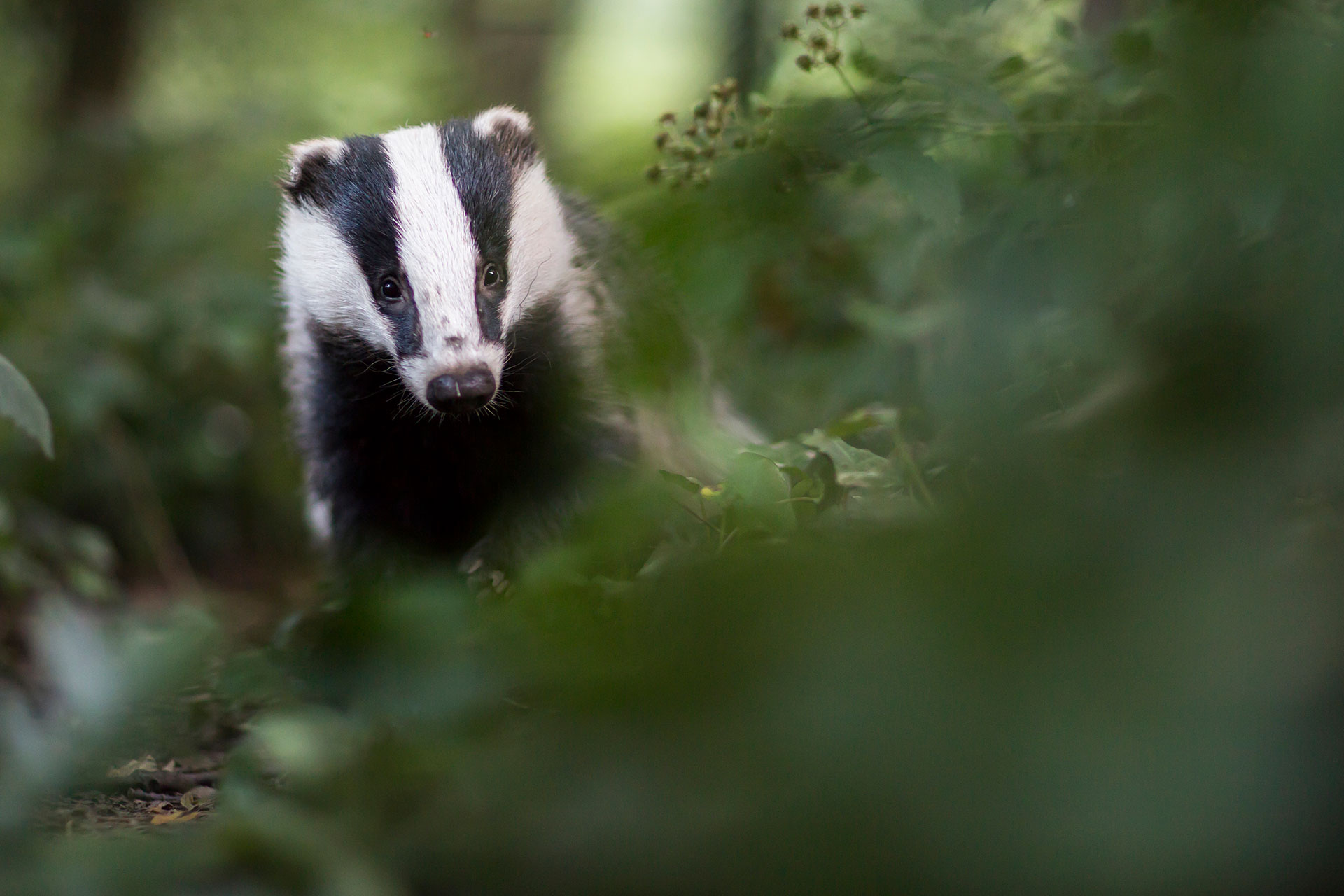

Individual variation in early-life telomere length and survival in a wild mammal (2019)
NBAF supported telomere length analysis using quantitative PCR methods.
Invisible
READ MORE
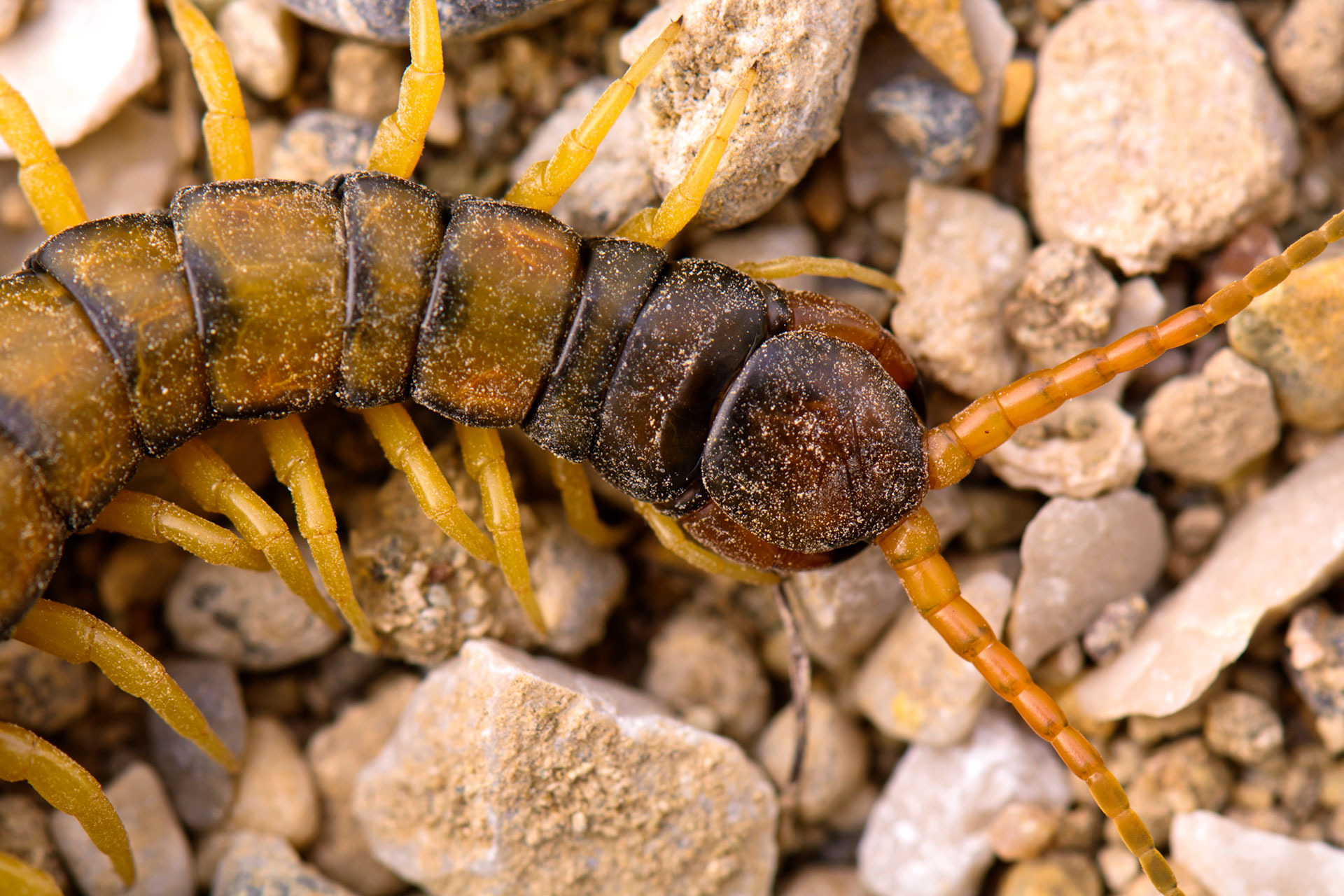

Parallel evolution of complex centipede venoms revealed by comparative proteotranscriptomic analyses (2019)
NBAF supported transcriptome sequencing.
Invisible
READ MORE
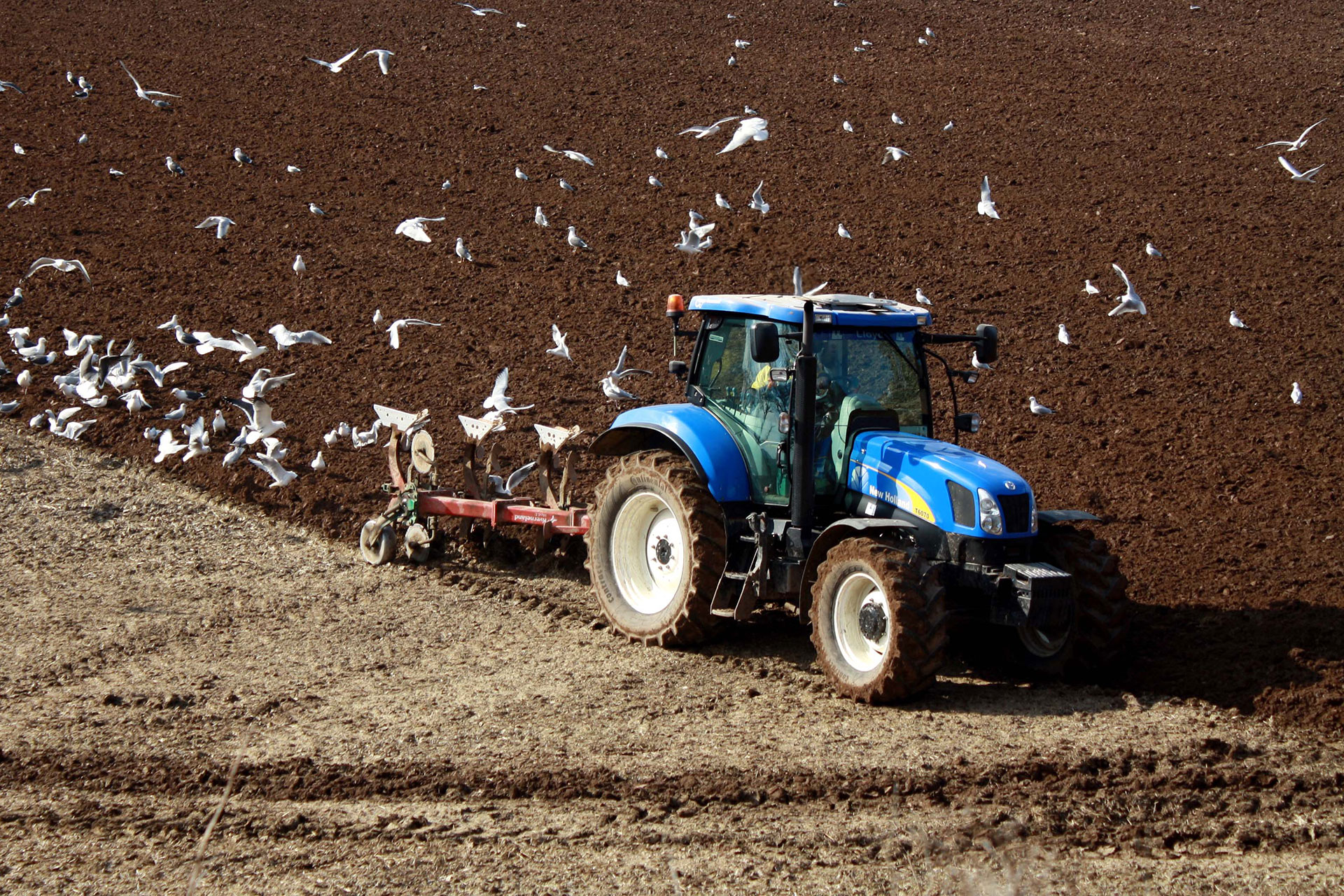

Divergent national-scale trends of microbial and animal biodiversity revealed across diverse temperate soil ecosystems (2019)
NBAF supported metabarcoding.
Invisible
READ MORE
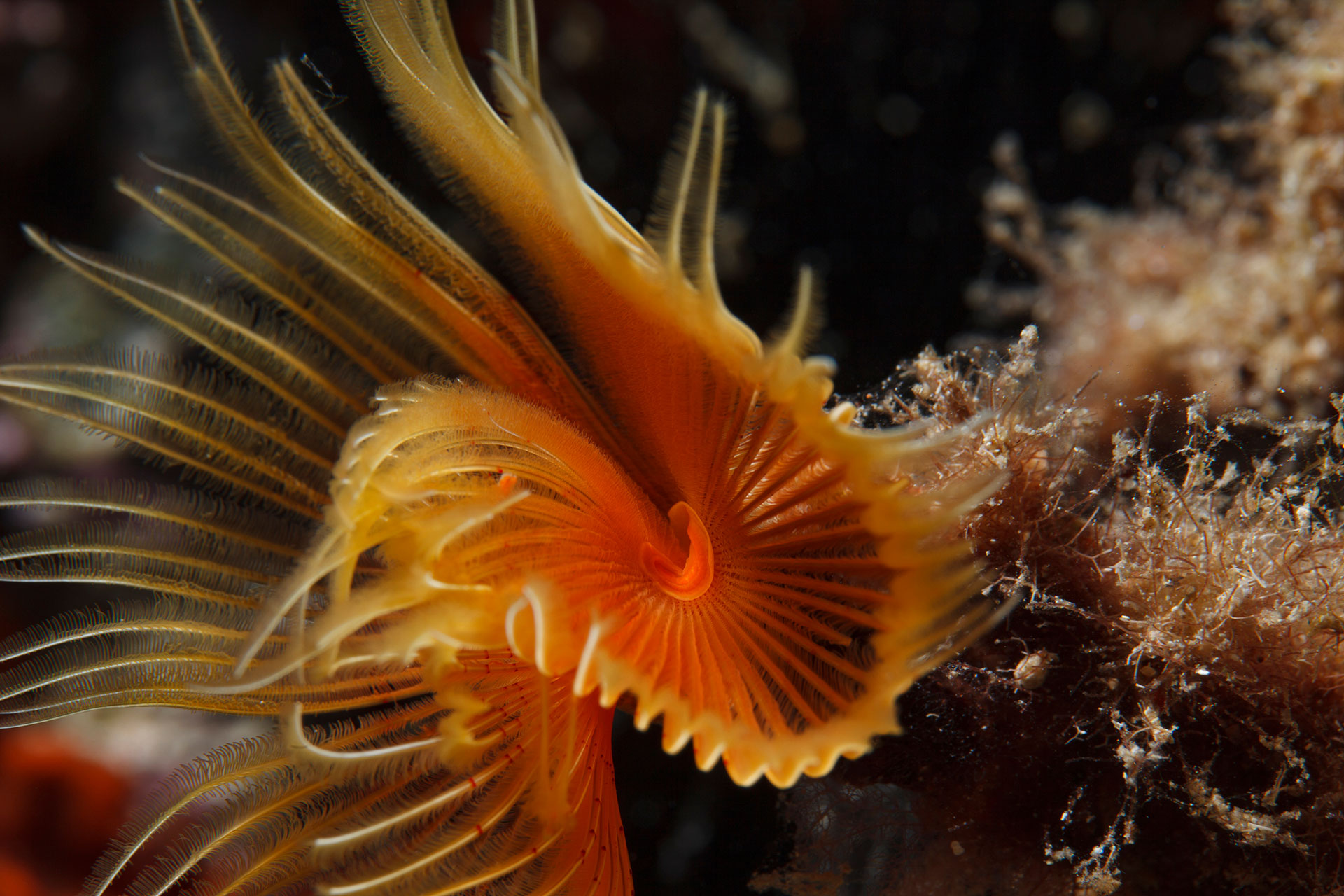

Lack of long-term acclimation in Antarctic encrusting species suggests vulnerability to warming (2019)
NBAF supported RNA-seq and metabarcoding.
Invisible
READ MORE
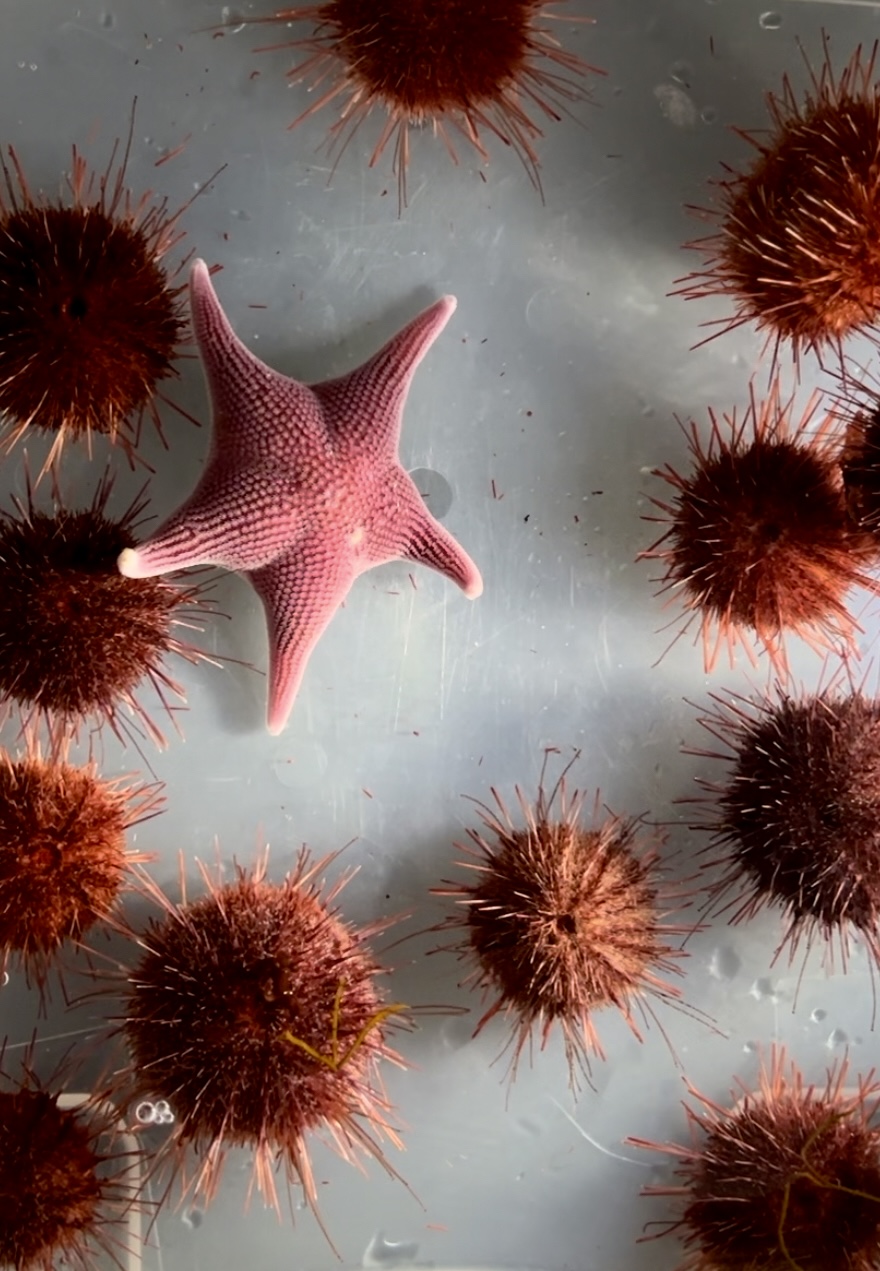
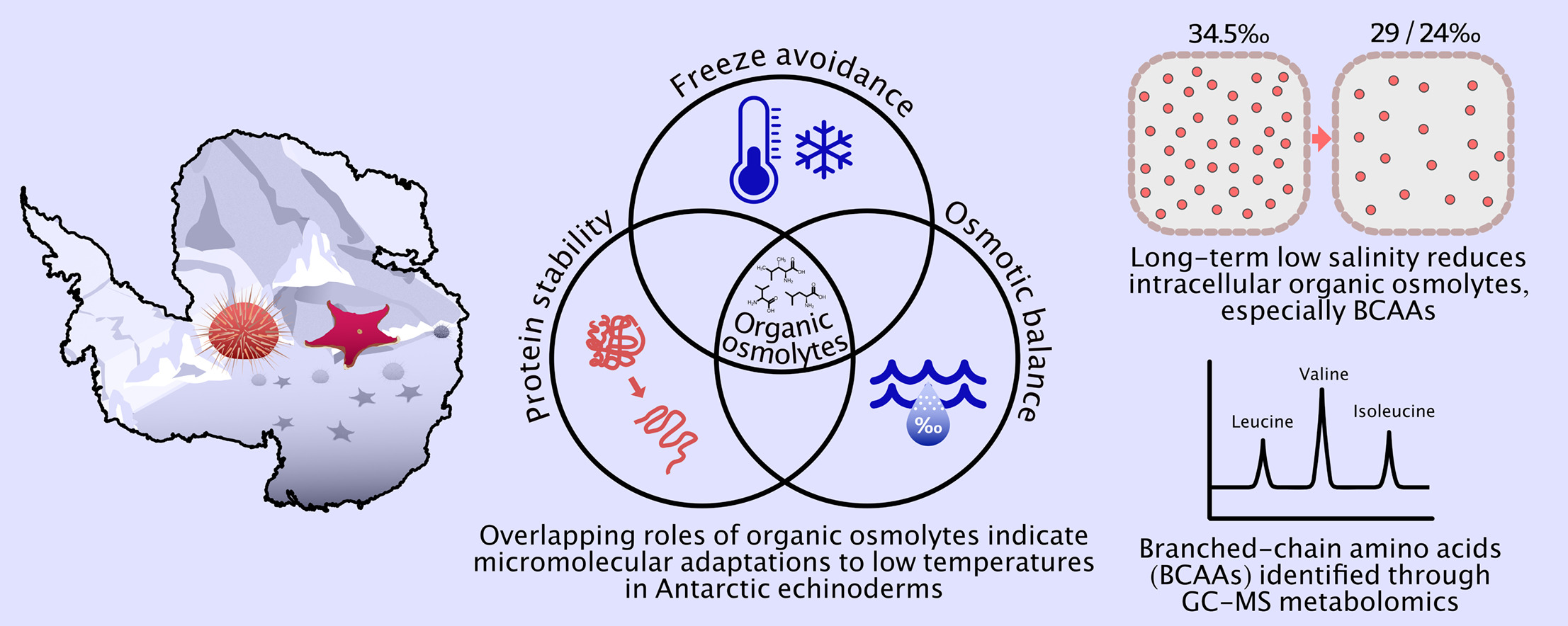

Metabolomics case study: Micromolecular adaptations in Antarctic echinoderms
Barrett NJ et al (2025), Science of The Total Environment 988, 179820. (10.1016/j.scitotenv.2025.179820)
Recipient of 2022 NEOF Independent Pilot Competition funding.
NEOF funding for this study enabled researchers at University of Cambridge and the British Antarctic Survey to work with CMR, leading to new insights into how protein stability and osmotic pressure is maintained in chronic low salinity and temperatures.
Invisible
READ MORE
For osmoconforming organisms, prolonged exposure to reduced salinity requires an adjustment to intracellular osmolyte levels to ensure an osmotic balance is maintained between the cell and external sea water. However, osmolytes—low-molecular-mass micromolecules—may also serve overlapping roles in freeze avoidance, desiccation resistance, and protein stabilisation. In Antarctic species living at or below 0 °C, multiple environmental stressors likely shape species-specific osmolyte profiles. Yet, the osmolytes utilised in osmotic acclimation and the broader micromolecular profile of Antarctic marine organisms remain poorly characterised. This study employed gas chromatography–mass spectrometry (GC–MS) to analyse the organic osmolyte composition of two endemic Antarctic echinoderms, the sea star Odontaster validus and sea urchin, Sterechinus neumayeri, following long-term acclimation (>12 weeks) to reduced salinity levels (29 ‰ and 24 ‰). Significant reductions in total tissue organic solute (osmolyte) concentrations after low salinity exposure indicated active cell volume regulation to reduce intracellular osmotic pressure. The osmolyte metabolic profiles of these Antarctic species appeared distinct from those of temperate echinoderms and other marine osmoconformers, suggesting a specialised adaptive response. Notably, the use of branched-chain amino acids (valine, leucine, and isoleucine) in cell volume regulation in O. validus, and to a lesser extent in S. neumayeri, suggests a micromolecular adaptation tailored to the extreme cold of the Antarctic environment.
Metabolomics case study: Using metabolomics to understand how environmental bacteria respond to pharmaceutical exposure
Mol. BioSyst. 2016, 12, 1367. https://doi.org/10.1039/c5mb00889a
PLoS ONE 2016 11: e0156509. https://doi.org/10.1371/journal.pone.0156509
Invisible
READ MORE
It has been known for decades that human pharmaceuticals are present in wastewater treatment plants and pollute rivers and estuaries at levels that affect aquatic organisms (see PNAS paper). What isn’t know is how microbial communities respond to exposure to APIs (active pharmaceutical ingredients) and this is likely to be at the phenotypic level where the microorganism responds to this stress. In addition, bacteria and fungi may transform APIs into other substances that could also affect both the planktonic and benthic communities.
In a paper published in Molecular BioSystems we used a combination of metabolic fingerprinting using Fourier transform infrared (FT-IR) spectroscopy with metabolic profiling using gas chromatography-mass spectrometry (GC-MS) to investigate phenotypic changes in an environmental isolate of Pseudomonas putida. Initially this bacterium was exposed to six APIs (acetaminophen, atenolol, diclofenac, ibuprofen, mefenamic acid, propranolol) at levels that did not affect the growth profile of the organism. Focussing on propranolol (a β-blocker used to treat hypertension) we found significant changes in carbon metabolism and in particular alterations in the concentration of energy related metabolites, including ATP. In addition, we found profound changes in the saturation levels of cardiolipins which may be related to the synthesis and activation of solvent extrusion pumps which would eliminate propranolol from inside these bacterial cells.
We went onto conduct an in-depth metabolomics study focussing on the efflux pumps involved in solvent extrusion which was published in PLoS ONE. In this study we included a series of efflux pump knockout mutants of P. putida DOT-T1E to assess the effect of this transporter on propranolol and metabolism.
These studies have opened up the exciting prospect of understanding how microbial communities adapt when exposed to toxic materials.
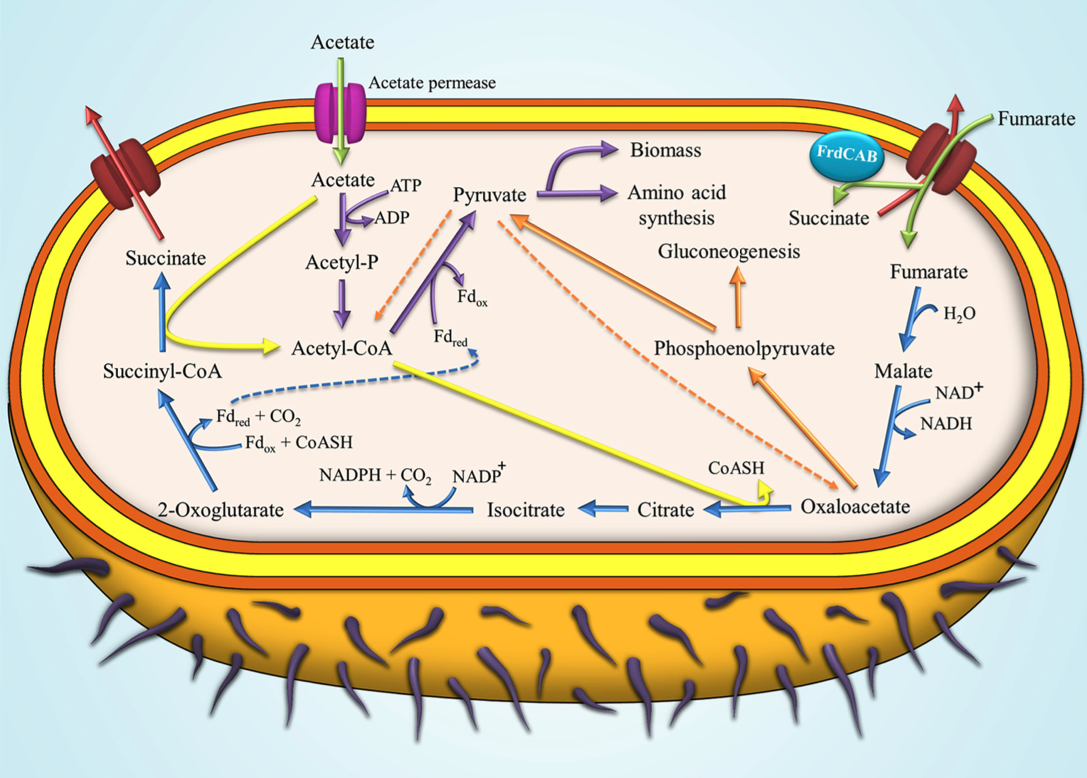
Metabolomics case study: Metabolic Profiling of Geobacter sulfurreducens during Industrial Bioprocess Scale-Up
Appl. Environ. Micro. 2015 81: 3288-3298. https://doi.org/10.1128/AEM.00294-15
Invisible
READ MORE
Geobacter sulfurreducens is an important bacterium as it is capable of global recycling of metals and therefore is important for bioremediation applications. In addition, this organism produces biogenic magnetite nanoparticles (BMNPs) which may have industrial application. However, during industrial scale-up of this BMNP production bioprocess we found an extended lag phase of 24 h for cells grown in 5-litre bioreactors compared to only 6 h those grown in 100 mL bottles.
In a paper published in Applied and Environmental Microbiology we investigated the phenotypic changes associated with the extended lag-phase during scale up of this bioprocess scale up using gas chromatography-mass spectrometry (GC-MS) for metabolic profiling.
GC-MS analysis showed that there were several changes in metabolites during scaleup and when relevant metabolites were overlaid onto a metabolic network of G. sulfurreducens we found that there was a limited availability of oxaloacetate and nicotinamide which would seem to be the main metabolic bottlenecks. Metabolite feeding experiments found that nicotinamide supplementation (1 mM) significantly reduced the lag phase in 5 L bioreactors from 24 h to 6 h, therefore resulting in a more rapid industrial bioprocess.
This study illustrates the exciting prospect of using metabolomics to enhance industrial scaleup of environmentally relevant microbial systems.
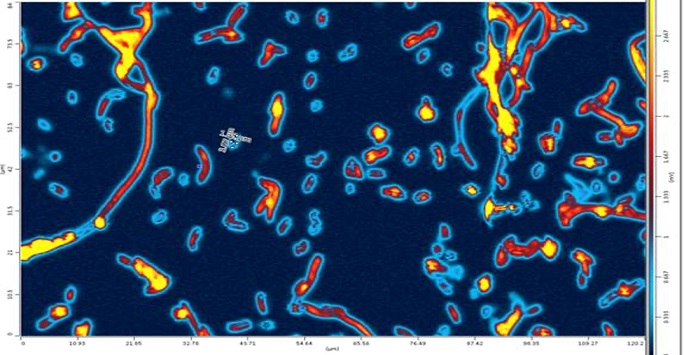
Metabolomics case study: Metabolism in Action: Developing stable isotope probing using Raman and infrared imaging for understanding metabolic flux at the single cell level
Anal. Chem. 2021, 93, 6, 3082–3088; https://doi.org/10.1021/acs.analchem.0c03967
Analyst 2021; 146, 1734-1746; https://doi.org/10.1039/D0AN02319A
Invisible
READ MORE
Complex microbial communities are found throughout ecosystems on the planet and these play essential functions that support animal and aquatic life.
Despite the vital role that bacteria play, linking these microbes with specific metabolic functions is difficult, since microbial communities consist of numerous and phylogenetically diverse microbes. This is compounded by the microscopic size of these microorganism (1-2 µm).
There is therefore a need to develop metabolomics solutions that generate images that are based on the biochemistry of the sample under analysis, and that provide information at micron and sub-micrometre scales. With the advent of spontaneous Raman and variants that can be tuned to specific vibrations (viz SRS and CARS), these, along with optical-photothermal infrared (O-PTIR) microscopy open up the tantalising opportunity to follow metabolism in microbial communities.

In a recent series of papers published in Analytical Chemistry and Analyst we have shown that the cellular uptake of stable isotope-labelled compounds by bacteria can be probed at the single-cell level using both Raman and infrared spectroscopies. This is due to the ability of these imaging techniques to monitor chemical vibrations that are affected by the incorporation of “heavy” atoms by cells and thus can be used to understand microbial systems. In this series of experiments, we have shown that it is possible to measure incorporation of 13C and 15N substrates, follow the dynamics of the rate of incorporation, as well as employing deuterated water to assess cross feeding in microbial communities.
This research has opened up the exciting prospect that these enabling chemical imaging technologies will guide the identification of primary substrate consumers in complex microbial communities in situ. This we believe represents a key step towards the characterisation of novel genes, enzymes and metabolic flux analysis in microbial consortia.
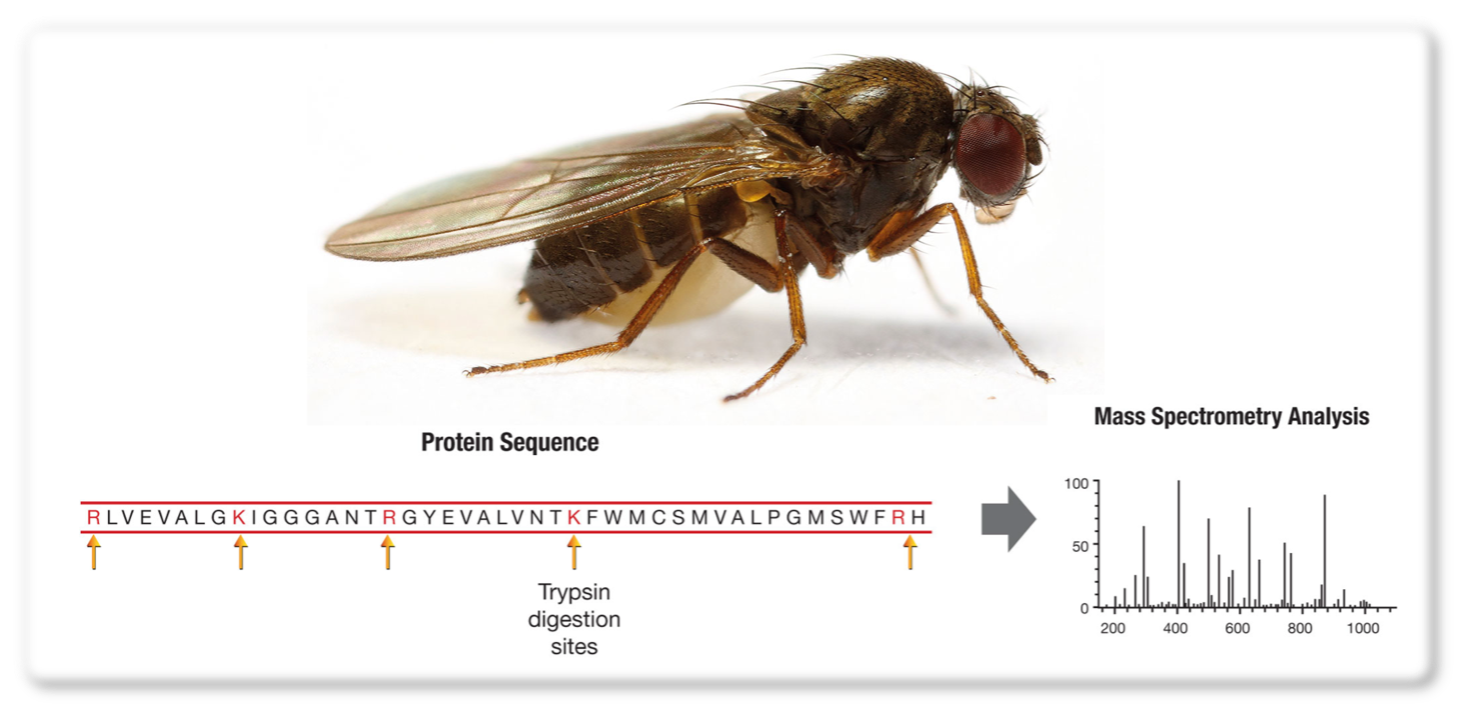

Proteomics case study: Targeting specific proteins or peptides
Aim: To identify sex peptides from Drosophila subobscura accessory gland samples.
Samples: 3 x 10 excised excretory glands from D. subobscura.
Results: There is no protein database for D. sub in UniProt, However, we were able to identify a suitable database by contacting other researchers who work with this organism. In addition, by pre-calculating the expected masses of trypsin cleavage products from the specific sex peptides of interest, we were able to perform target-ed analysis of these peptides to get a positive identification.
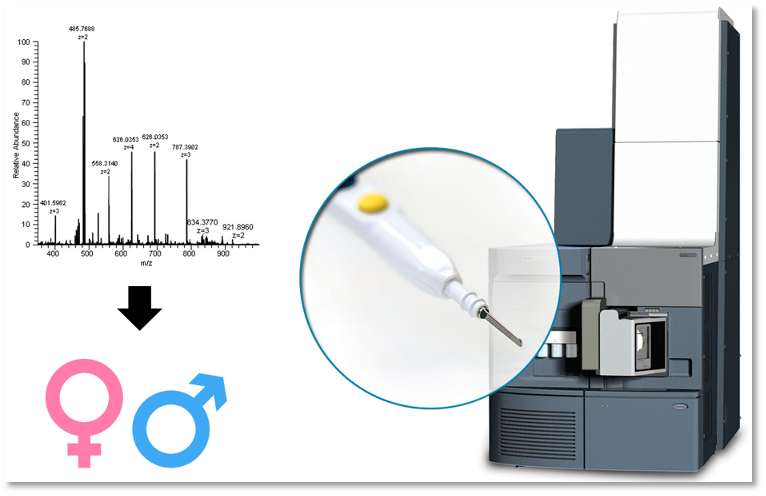
Proteomics case study: Species identification
Aim: Rapid identification of species and sex, based on pattern recognition of mass spectra, for closely related species that are notoriously difficult to identify by eye.
Samples: 150-170 whole mosquito samples of each of D. bifasciata, D. pseudoobscura, D. suboscura, D. melanogaster and D. simulans were used to build a species model; 400 male and 400 female mosquitoes across the 5 species were used to build the sex determination model.
Results: 95 % accuracy in species determination, and 99 % accuracy in sex determination.
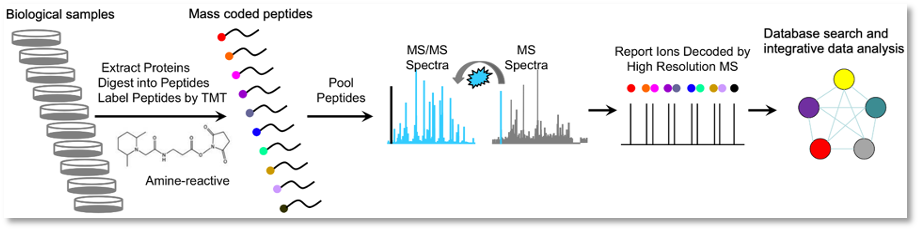
Proteomics case study: Impact of viral infection on host proteome
Aim: To quantify changes to the Asian tiger mosquito proteome when exposed to virus.
Samples: 3 replicates of 14 conditions.
Results: By using TMT isobaric labelling, more than 5000 proteins were quantified including 11 out of 12 virus proteins. Proteins were identified from a genome database.
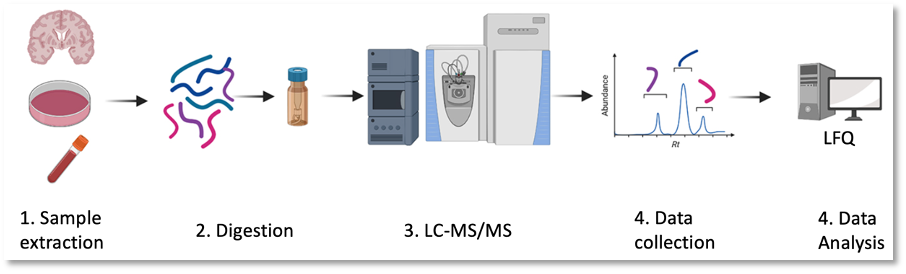
Proteomics case study: Secretome proteome of pseudomonas aeruginosa
Aim: To identify the difference in multi-drug resistance strain verse the wild type.
Samples: 5 replicates of 2 pseudomonas aeruginosa strains.
Results: Proteins were concentrated from 10 mL to 100 μL by strataclean beads. Digestion was conducted from straight from the strataclean beads. Over 1000 proteins were identified in each sample. Samples were processed for label free quantification.
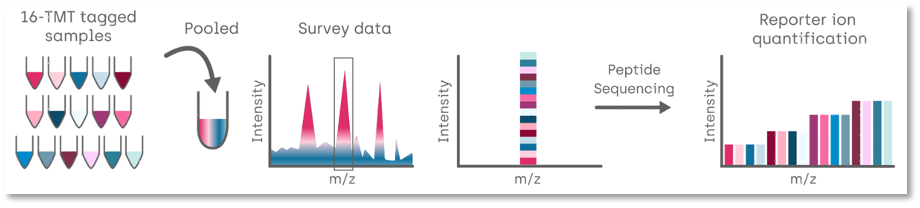
Proteomics case study: Identification of phosphorylation targets for a phosphatase in human cell line
Aim: To determine which proteins are phosphorylated and the down stream effects of phosphorylation.
Samples: 4 replicates of 4 clonal cell lines.
Results: Samples were labelled with TMT isobaric labelling. The sample was fractionated into 12 fractions and then each used to deter mine i) global protein changes and ii) phosphopeptide changes. Around 6000 proteins and in excess of 10,000 phosphopeptides were identified, with the vast majority being quantified across all 16 samples.
